 Both minima and saddle points are stationary points on the energy surface, wherever the primary derivative of the energy function is zero with regard to all the coordinates. The highest point on the pathway between two minima is of particular interest and is understood as the saddle point, with the arrangement of the atoms being the transition structure. To spot those geometries of the system that correspond to minimum points on the energy surface we tend to use a Minimization algorithm. The minimum with the very lowest energy is known as the global energy minimum. There is also a really sizable amount of minima on the energy surface. Minimum energy arrangements of the atoms correspond to stable states of the system any movement off from a minimum provides a configuration with a better energy. 
In molecular modeling we tend to are particularly curious about minimum points on the energy surface. Finally, the calculation should be analyzed, not solely to calculate properties however additionally to see that it’s been performed properly. The second stage of a molecular modeling study is that the calculation itself, such as an energy Minimization, a molecular dynamics or Monte Carlo simulation, or a Conformations search. These models enable the energy of any arrangement of the atoms and molecules within the system to be calculated and permit the modeler to work out how the energy of the system varies because the positions of the atoms and molecules change. The two most typical models that are utilized in molecular modeling are quantum mechanics and molecular mechanics. Within the initial stage a model is chosen to explain the intra- and inter-molecular interaction within the system. Most molecular modeling studies involve three stages. Molecular simulation techniques need specific extra computational and code software system. The fundamental computational technique to perform molecular modeling is simulation. The applying fields of molecular modeling regard computational chemistry, drug design, computational biology and materials science. Molecular modeling relies on the event of theoretical and computational methodologies, to model and study the behavior of molecules, from little chemical systems to big biological molecules and material assemblies. An efficient optimization protocol could be devised from these methods in conjunction with a larger space exploration algorithm, e.g. Although energy Minimization is a tool to achieve the nearest local minima, it is also an indispensable tool in correcting structural anomalies, viz. 
Three common algorithms used for this optimization are steepest descent, conjugate gradient and Newton–Raphson. Various algorithms have been formulated by varying the use of derivatives. The goal of energy Minimization is to find a set of coordinates representing the minimum energy conformation for the given structure. All these quantitative terms have been parameterized and are collectively referred to as the ‘force-field’, for e.g. the long-range forces between charged and partially charged atoms. Electrostatic energy accounting for the Coulomb’s Law m protein structure, i.e. A van der Waals term (also called Leonard-Jones potential) to ensure that atoms do not have steric clashes. Dihedral energy, due to the dihedral angles. Bond energy and angle energy, representative of the covalent bonds, bond angles. 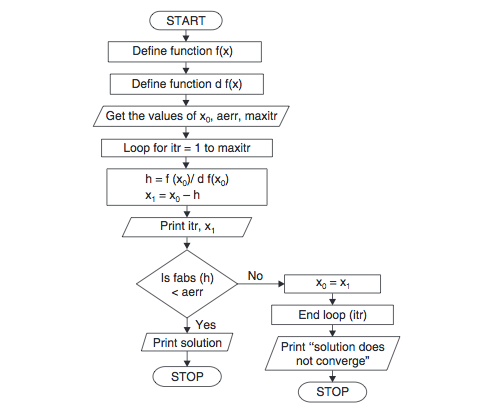
This energy function consists of several components: 1. 
The energy of a protein can be defined as a function of its atomic coordinates. The energetic state of a protein is one of the most important representative parameters of its stability. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
